Peptide Library Synthesis | Omizzur

Peptide Library Synthesis is an important tool in research for bioactivity-screening A peptide library is a collection of small peptides of a certain length. The establishment of peptide library is based on two facts:1.small peptides with pivotal residues can simulate antigenic determinants on proteins; 2.the interaction and recognition between proteins are realized through the interaction between local peptide fragments.By screening hundreds of peptide combinations scientists can do lots of bioactive studies:epitope peptide mapping, drug target validation ,T cell immunotherapy and the vaccine development so on.
A typical peptide library contains of 20-25 amino acids,With OMIZZUTM peptide library technology we can synthesize peptides of up to 30 amino acids in length in individually vials .Each peptide follow strict quality system to ensure accuracy and prevent any cross contaminants.With our high throughput peptide technology, Omizzur provide comprehensive options for peptide library design and peptide purity ranging from crude to 99% for fullest QC services .
Comprehensive Peptide Library Design Tools
A wide range of library design tools:Overlapping, Truncated,T cell truncated,Alanine scan,Random library,Scrambled,Positional scan,Conbinatorial 2/3-Positional Scan to meet different demands.
Avoid Cross Contamination
Peptides are lyophilized in 96 tube-plates or individual vials to avoid any cross contamination.
Overall Peptide Modifications
Omizzur incorporate all round peptide modifications: N/C-terminals modification,special amino acids,peptide conjugates, biotin, phosphorylation fluorescence labeling so on.
Stringent Quality Control System
Custom peptide library follows strict Omizzur Quality System,By automatic OMIZZUTM Synthesis platform,Omizzur ensure to provide high purity, no batch difference for each peptide products.
| Peptide library | Crude I | Purified I | Purified II |
| Length | 5-30AA | 5-15AA | 16-30AA |
| Purity | Crude | >70% | >98% |
| Quantity | 3-20mg | 3-9mg | 3-9mg |
| Deliver | 7-15 days | 10-15 days | 15-20days |
| Min order | 24 peptides | 24 peptides | 24 peptides |
| Peptide modifications | N/C-terminals modification,special amino acids,peptide conjugates, biotin, phosphorylation etc | ||
| Analysis report | COA | HPLC+COA+MS | HPLC+COA+MS |
| Applications | Library types | Purity suggestion |
| •Antigen epitope mapping. •T-cell epitope identification •Protein sequence scanning | Overlapping Library | Crude |
| •T-cell Immunotherapy •Metabolism study of peptide drugs | Truncation Library | Crude or ≥70% |
| •Antibody epitope mapping •Enzyme substrates study | Combinatorial Positioning Scanning Library Overlapping Library | Crude or ≥70% |
| •Biological assays | Scrambled Peptide Library Overlapping Library | Up to 70% |
| •Vaccine development | Alanine Scanning Library | Up to 90% |
Peptide library is a mixture of peptide fragment of certain length, which contains all possible arrangement information of amino acids .There are two kinds of peptide library, one is synthetic peptide library, the other is phage peptide library.The fragment in the peptide library has the property of binding to the corresponding specific protein,A single specific fragment can be selected from the peptide library by using the labeled specific protein,The markers which used for labeling proteins can be enzymes, fluorescein, biotin, isotopes, We call this labeled specific protein as probe.
Using monoclonal antibody as probe can screening peptide fragment with high affinity from the synthetic peptide library.This specific epitope information can provide important inspiration for the development of peptide vaccine;Use cytokine as probe to screening high affinity peptide fragment from peptide library may be the similar structure of receptor cell.Based on this information, scientist can develop specific synthetic peptide antagonists for cytokine receptor, which may be developed into a peptide drug.
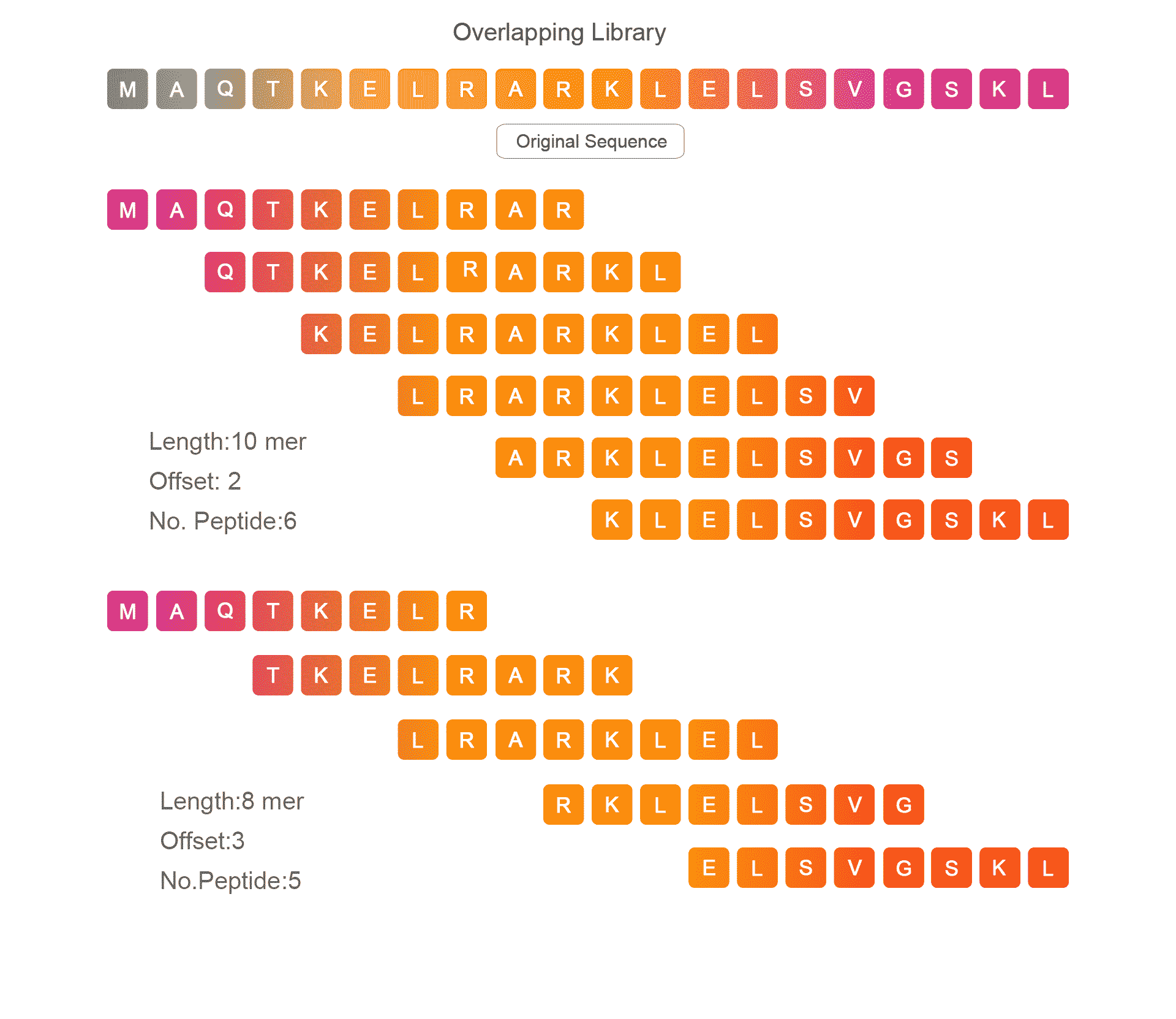
Overlapping Peptide Library
It is mainly used for epitope mapping of antigen and protein scanning.by this method we can find out which amino acids fragment that divided from peptide or protein can contribute the biological activity.Overlapping Library is mainly determined by two key parameters: peptide length and offset number.The length of peptide is generally 5-15 AA , and the number of offset number is generally 2-5.
The T cell epitopes are generally linear short peptides in holoprotein, so overlapping library is also a common method to screen T cell epitopes.The high-throughput AurPepTM synthesis technology provides the possibility for this research work.
Application area: Antigen epitope mapping ,Holoprotein scanning,T-cell epitope identification.

Truncation Peptide Library
Identification of shortest amino acid sequences which related to biological activity,by systematically removing the amino acids on both sides of the original peptide sequence can identify the minimum range of antigen epitopes.
Application area: Shortest bioactive sequence identification,Metabolism of peptide drugs in vivo.
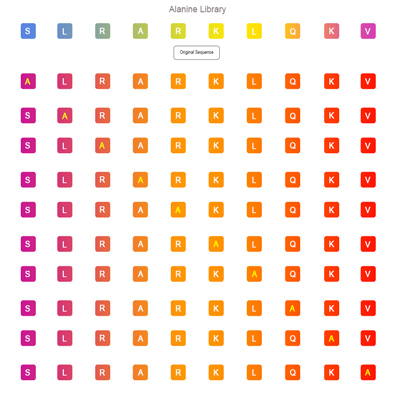

Alanine Peptide Library
Alanine is the smallest chiral amino acid In 20 kinds of natural amino acids. It can be used to identify the effect of each site of the target peptide on its biological activity by individually replacing each amino acid in sequence .When the essential amino acid residues are replaced by alanine, the peptides will correspondingly reduce the biological activity.
Application area:Study the effect of a specific amino acid on the whole protein function.
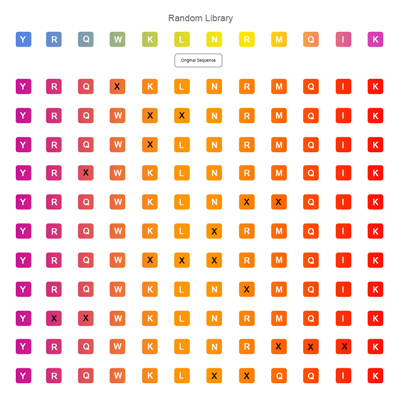
Random Peptide Library
It uses 20 kinds of natural amino acids to replace the selected amino acid residues in the original peptide simultaneously and randomly by shotgun approach, so as to form as many random mutations as possible.In this way, peptides with specific functions or activities can be identified preliminarily in a group of active sequences.
Application area:Optimization peptide sequence ,Drug discovery
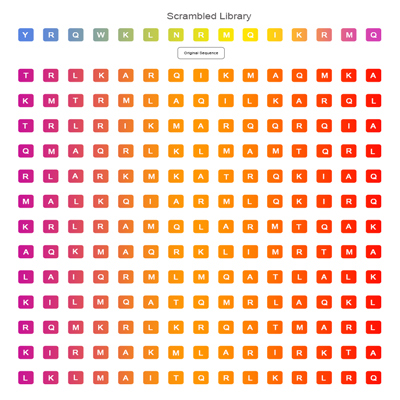

Scrambled Peptide Library
It is mainly used for peptide screening or peptide sequence optimization which targeting as protein, antibody or DNA.The design of this library is based on the internal replacement of the original peptide sequence. It is the most mutated peptide library ,also a random screening tool for finding new targets.
Application area:Peptide screening,Peptide sequence optimization
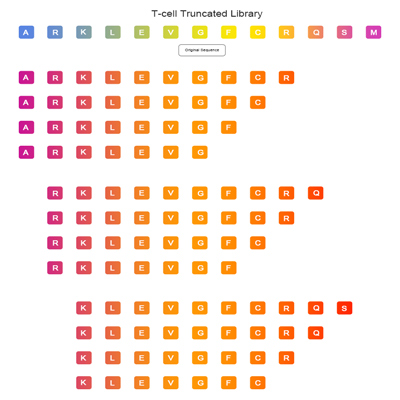

T-cell Truncated Library
T-cell Truncated Library allows high-resolution identification of T cell epitopes across a protein.The peptide was gradually cut from both ends to form peptide library, and then the peptide library was mixed into peptide pool to screen the best epitope of T cell. T cell truncated peptide library provides a powerful guarantee for screening the best epitope of T cell of interest.
Application area: Test all possible T-cell epitopes in protein
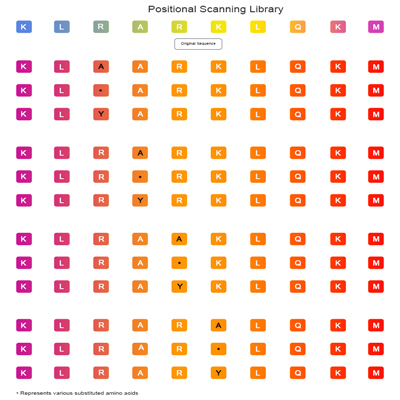

Combinatorial Positional Scanning Peptide Library
Combinatorial Positional scanning library is an important tool for optimization of peptide sequences.and is an upgrade to positional scanning.After screening out the important sites affecting activity by alanine library,the modification schemes to improve the activity of peptides can be quickly discovered by varying systematically 2 or more positions at a time(replace each site of the peptide with unnatural/natural amino acids,D-amino acids,Phe/Tyr derivatized amino acids,Phosphorylation/N-Methylation etc)

It will be 20×20 = 400 peptides to be produced in the process of 2-Positional Scanning with 20 kinds of natural amino acids, 3-Positional Scanning would be 8000 peptides total.Due to the huge work of amino acid substitution, our self developed AuaPepTM Synthesis platform can help you to quickly generate this matrix combination. Please tell us the sequence, the site which to be studied, the purity and quantity, our synthesis platform will automatically help you to generate this matrix.
Application area: Improvement for peptide biological activity.,Study on the activity of enzyme substrates.

Omizzur has introduced high-throughput automatic peptide synthesis equipment with independently developed OMIZZUTM synthesis platform to produce mg level and multi sequence peptide products to meet the peptide library demand of clients. Omizzur ’s software will automatically help you build peptide library when you let us the know the sequence, purity and other related requirements you want to study,and the results will be sent back to you by email together with the quotation for your references.
References
1.Smith G P. Surface Display and Peptide Libraries. Gene,1993,128.1-2
2.Devlin JJ,Pangniban L C ,Devlin P E. Random Peptide Libraries:A Sourse of Specific Protein Binding Molecules Science,1990,249:404-406
3.Scott JK,Smith G P.Searching for Peptide Ligands with an Epitope Library.Science,1990,249:386-390
4.Houghten RA,Pinilla C,Blondelle SE,et al,Generation and use of synthetic peptide combinatorial libraries for basic research and drug discovery [J].Nature,1991,354:84-86
5.Ohlmeyer MH,Swanson RN,Dillard LW,et al,Complex synthetic chemical libraries indexed with molecular tags[J],Proc Natl Acad Sci,1993,90:10922-10926
Customer Service Center
Soild-Phase Peptide Synthesis
Copyright © 2020 Omizzur Inc | Terms & Conditions | Privacy Notice | Sitemap